Scaling
Compute Scaling Relationships for Malaria Metrics
January 28, 2026
Source:vignettes/Scaling.Rmd
Scaling.Rmdramp.work has functions to compute
scaling relationships: how do various measures of malaria change across
the spectrum of malaria transmission intensity?
This vignette explains the functions in
ramp.work that:
compute scaling relationships
visualize scaling relationships
A longer introduction to scaling can be found in SimBA.
xds_scaling enables Metric
Conversion
Pseudocode
Suppose that we have defined a model family: we modify a single parameter that changes the mean intensity of exposure or transmission, but we leave the rest of the parameters fixed.
To study the relationship between exposure and malaria outcomes, we might vary the mean annual EIR.
To study the relationship between mosquito density and outcomes, we might vary the average annual emergence rate. \(\Lambda.\)
To compute scaling relationships in systems with a seasonal forcing pattern, we will need to compute the stable orbits and link the outputs to the average annual values.
The function xds_scaling
handles this task.
-
create a mesh of values for mean forcing, either
the mean daily EIR, \(\bar E.\)
the mean daily emergence rate for adult mosquitoes, \(\bar \Lambda\) or
Lambda.
-
for each mesh value:
compute and store the stable orbits for important terms
compute and store the mean values of the stable orbits
Example
Load the required packages:
Set up a model forced by the EIR with a seasonal pattern, and we use
ramp.xds::show_season to see the pattern:
Spar <- makepar_F_sin(bottom = 0.2, pw=2)
model <- xds_setup_eir(eir = 1/365, season_par = Spar)
model <- xds_solve(model)
show_season(model)
The function xds_scaling
computes and stores the values:
xds_scaling$scalingstores the average annual values by namexds_scaling$scaling$stable_orbitsstores the stable orbits
xds_scaling(model) -> model
names(model$scaling)## [1] "aeir" "eir" "pr" "ni"
## [5] "stable_orbits"A plot_eirpr plot the PfPR as a function of the
average annual PfEIR on a semi-log plot. A function
add_eirpr_orbits adds the stable orbits for an indexed
subset. We use the virisLite::turbo color scheme so it is
easy to relate the orbit and the mean value.
library(viridisLite)
xds_plot_eirpr(model)
ix_subset = c(9, 13, 17, 21)
add_eirpr_orbits(ix_subset, model, clrs = turbo(25))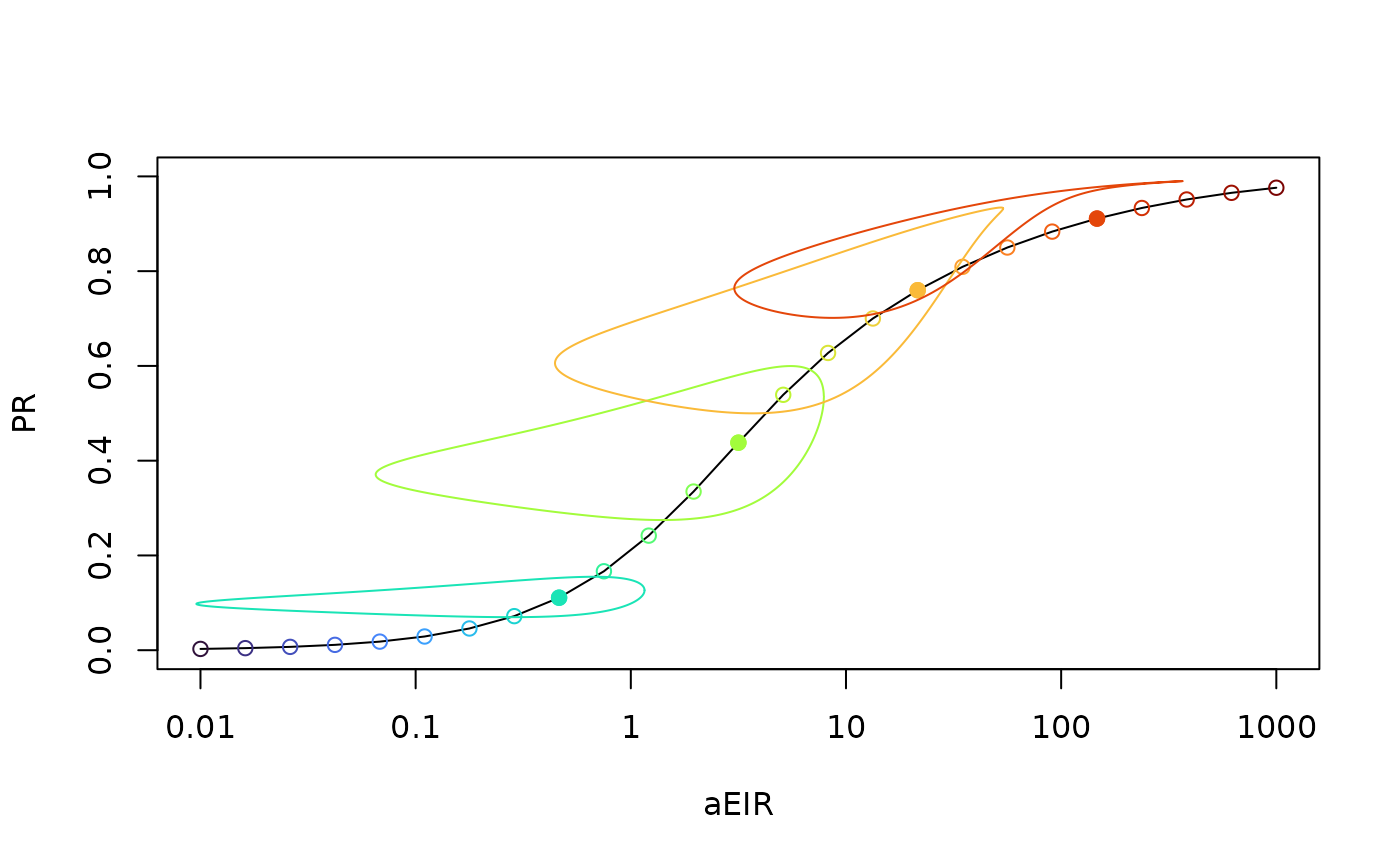
Notes
xds_scalingdispatches onxds_obj$forced_by,which is set up either byxds_setup_eiror by themake_L_obj_trivial.The function
xds_scaling.eir()creates a mesh over logged values of on an even mesh for values oflog(aEIR)running from \(10^{-2}\) up to \(10^{3}.\)The function
xds_scaling.Lambda()computes a creates a crude mesh. The first step is to compute an approximate pseudo-threshold value \(L.\) The initial mesh looks has five values \(c(L/100, L/5, L, 5L, 100L).\) Subsequent values are identified by picking a new value in the interval that has the largest change in PfPR. After adding the new value, the